Knowledge-primed Neural Networks (KPNNs) for single cell data#
In this tutorial we will show how CORNETO can be used to build custom neural network architectures informed by prior knowledge. We will see how to implement a knowledge-primed neural network1. We will use the single cell data from the publication “Knowledge-primed neural networks enable biologically interpretable deep learning on single-cell sequencing data”, from Nikolaus Fortelny & Christoph Bock, where they used a single-cell RNA-seq dataset they previously generated2, which measures cellular responses to T cell receptor (TCR) stimulation in a standardized in vitro model. The dataset was chosen due to the TCR signaling pathway’s complexity and its well-characterized role in orchestrating transcriptional responses to antigen detection in T cells.
Why CORNETO?#
In the original publication, authors built a KPNN by searching on databases, building a Direct Acyclic Graph (DAG) by running shortest paths from TCR receptor to genes. However, this approach is not optimal. CORNETO, thanks to its advanced capabilities for modeling and optimization on networks, provides methods to automatically find DAG architectures in an optimal way.
In addition to this, CORNETO provides methods to build DAG NN architectures with ease using Keras +3, making KPNN implementation very flexible and interoperable with backends like Pytorch, Tensorflow and JAX.
How does it work?#
Thanks to CORNETO’s building blocks for optimization over networks, we can easily model optimization problems to find DAG architectures from a Prior Knowledge Network. After we have the backbone, we can convert it to a neural network using the utility functions included in CORNETO.
References#
Fortelny, N., & Bock, C. (2020). Knowledge-primed neural networks enable biologically interpretable deep learning on single-cell sequencing data. Genome biology, 21, 1-36.
Datlinger, P., Rendeiro, A. F., Schmidl, C., Krausgruber, T., Traxler, P., Klughammer, J., … & Bock, C. (2017). Pooled CRISPR screening with single-cell transcriptome readout. Nature methods, 14(3), 297-301.
Download and import the single cell dataset#
import os
import tempfile
import urllib.parse
import urllib.request
import numpy as np
import pandas as pd
import scanpy as sc
import corneto as cn
with urllib.request.urlopen("http://kpnn.computational-epigenetics.org/") as response:
web_input = response.geturl()
print("Effective URL:", web_input)
files = ["TCR_Edgelist.csv", "TCR_ClassLabels.csv", "TCR_Data.h5"]
temp_dir = tempfile.mkdtemp()
# Download files
file_paths = []
for file in files:
url = urllib.parse.urljoin(web_input, file)
output_path = os.path.join(temp_dir, file)
print(f"Downloading {url} to {output_path}")
try:
with urllib.request.urlopen(url) as response:
with open(output_path, "wb") as f:
f.write(response.read())
file_paths.append(output_path)
except Exception as e:
print(f"Failed to download {url}: {e}")
print("Downloaded files:")
for path in file_paths:
print(path)
Effective URL: https://medical-epigenomics.org/papers/fortelny2019/
Downloading https://medical-epigenomics.org/papers/fortelny2019/TCR_Edgelist.csv to /var/folders/b4/gwkwsdb93sv11rtztqbm3l040000gn/T/tmpgy9f05hv/TCR_Edgelist.csv
Downloading https://medical-epigenomics.org/papers/fortelny2019/TCR_ClassLabels.csv to /var/folders/b4/gwkwsdb93sv11rtztqbm3l040000gn/T/tmpgy9f05hv/TCR_ClassLabels.csv
Downloading https://medical-epigenomics.org/papers/fortelny2019/TCR_Data.h5 to /var/folders/b4/gwkwsdb93sv11rtztqbm3l040000gn/T/tmpgy9f05hv/TCR_Data.h5
Downloaded files:
/var/folders/b4/gwkwsdb93sv11rtztqbm3l040000gn/T/tmpgy9f05hv/TCR_Edgelist.csv
/var/folders/b4/gwkwsdb93sv11rtztqbm3l040000gn/T/tmpgy9f05hv/TCR_ClassLabels.csv
/var/folders/b4/gwkwsdb93sv11rtztqbm3l040000gn/T/tmpgy9f05hv/TCR_Data.h5
# The data contains also the original network they built with shortest paths.
# We will use it to replicate the study
df_edges = pd.read_csv(file_paths[0])
df_labels = pd.read_csv(file_paths[1])
# Import the 10x data with Scanpy
adata = sc.read_10x_h5(file_paths[2])
df_labels
| barcode | TCR | |
|---|---|---|
| 0 | AAACCTGCACACATGT-1 | 0 |
| 1 | AAACCTGCACGTCTCT-1 | 0 |
| 2 | AAACCTGTCAATACCG-1 | 0 |
| 3 | AAACCTGTCGTGGTCG-1 | 0 |
| 4 | AAACGGGTCTGAGTGT-1 | 0 |
| ... | ... | ... |
| 1730 | TTTCCTCGTCATGCCG-2 | 1 |
| 1731 | TTTGCGCGTAGCCTCG-2 | 1 |
| 1732 | TTTGGTTAGATACACA-2 | 1 |
| 1733 | TTTGGTTGTATGAATG-2 | 1 |
| 1734 | TTTGGTTTCCAAGTAC-2 | 1 |
1735 rows × 2 columns
df_edges
| parent | child | |
|---|---|---|
| 0 | TCR | ZAP70 |
| 1 | ZAP70 | MAPK14 |
| 2 | MAPK14 | FOXO3 |
| 3 | MAPK14 | STAT1 |
| 4 | MAPK14 | STAT3 |
| ... | ... | ... |
| 27574 | HMGA1 | MTRNR2L9_gene |
| 27575 | MYB | C12orf50_gene |
| 27576 | MYB | TRPC5OS_gene |
| 27577 | SOX2 | TRPC5OS_gene |
| 27578 | CRTC1 | MTRNR2L9_gene |
27579 rows × 2 columns
adata.var
| gene_ids | |
|---|---|
| DDX11L1 | ENSG00000223972 |
| WASH7P | ENSG00000227232 |
| MIR6859-2 | ENSG00000278267 |
| MIR1302-10 | ENSG00000243485 |
| MIR1302-11 | ENSG00000274890 |
| ... | ... |
| Tcrlibrary_RUNX2_3_gene | Tcrlibrary_RUNX2_3_gene |
| Tcrlibrary_ZAP70_1_gene | Tcrlibrary_ZAP70_1_gene |
| Tcrlibrary_ZAP70_2_gene | Tcrlibrary_ZAP70_2_gene |
| Tcrlibrary_ZAP70_3_gene | Tcrlibrary_ZAP70_3_gene |
| Cas9_blast_gene | Cas9_blast_gene |
64370 rows × 1 columns
# We can normalize the data, however, it is better to avoid
# preprocessing the whole dataset before splitting in training and test
# to avoid data leakage.
# NOTE: Normalization can be done inside the cross-val loop
# sc.pp.normalize_total(adata, target_sum=1e6)
# Log-transform the data does not leak data as it does not estimate anything
sc.pp.log1p(adata)
adata.obs
| AAACCTGAGAAACCAT-1 |
|---|
| AAACCTGAGAAACCGC-1 |
| AAACCTGAGAAACCTA-1 |
| AAACCTGAGAAACGAG-1 |
| AAACCTGAGAAACGCC-1 |
| ... |
| TTTGTCATCTTTACAC-2 |
| TTTGTCATCTTTACGT-2 |
| TTTGTCATCTTTAGGG-2 |
| TTTGTCATCTTTAGTC-2 |
| TTTGTCATCTTTCCTC-2 |
1474560 rows × 0 columns
barcodes = adata.obs_names
barcodes
Index(['AAACCTGAGAAACCAT-1', 'AAACCTGAGAAACCGC-1', 'AAACCTGAGAAACCTA-1',
'AAACCTGAGAAACGAG-1', 'AAACCTGAGAAACGCC-1', 'AAACCTGAGAAAGTGG-1',
'AAACCTGAGAACAACT-1', 'AAACCTGAGAACAATC-1', 'AAACCTGAGAACTCGG-1',
'AAACCTGAGAACTGTA-1',
...
'TTTGTCATCTTGGGTA-2', 'TTTGTCATCTTGTACT-2', 'TTTGTCATCTTGTATC-2',
'TTTGTCATCTTGTCAT-2', 'TTTGTCATCTTGTTTG-2', 'TTTGTCATCTTTACAC-2',
'TTTGTCATCTTTACGT-2', 'TTTGTCATCTTTAGGG-2', 'TTTGTCATCTTTAGTC-2',
'TTTGTCATCTTTCCTC-2'],
dtype='object', length=1474560)
gene_names = adata.var.index
print(gene_names)
Index(['DDX11L1', 'WASH7P', 'MIR6859-2', 'MIR1302-10', 'MIR1302-11', 'FAM138A',
'OR4G4P', 'OR4G11P', 'OR4F5', 'RP11-34P13.7',
...
'Tcrlibrary_RUNX1_1_gene', 'Tcrlibrary_RUNX1_2_gene',
'Tcrlibrary_RUNX1_3_gene', 'Tcrlibrary_RUNX2_1_gene',
'Tcrlibrary_RUNX2_2_gene', 'Tcrlibrary_RUNX2_3_gene',
'Tcrlibrary_ZAP70_1_gene', 'Tcrlibrary_ZAP70_2_gene',
'Tcrlibrary_ZAP70_3_gene', 'Cas9_blast_gene'],
dtype='object', length=64370)
len(set(df_labels.barcode.tolist()))
1735
len(set(barcodes.tolist()))
1474560
matched_barcodes = sorted(set(barcodes.tolist()) & set(df_labels.barcode.tolist()))
len(matched_barcodes)
1735
# This is the InPathsY data in the original code of KPNNs
df_labels
| barcode | TCR | |
|---|---|---|
| 0 | AAACCTGCACACATGT-1 | 0 |
| 1 | AAACCTGCACGTCTCT-1 | 0 |
| 2 | AAACCTGTCAATACCG-1 | 0 |
| 3 | AAACCTGTCGTGGTCG-1 | 0 |
| 4 | AAACGGGTCTGAGTGT-1 | 0 |
| ... | ... | ... |
| 1730 | TTTCCTCGTCATGCCG-2 | 1 |
| 1731 | TTTGCGCGTAGCCTCG-2 | 1 |
| 1732 | TTTGGTTAGATACACA-2 | 1 |
| 1733 | TTTGGTTGTATGAATG-2 | 1 |
| 1734 | TTTGGTTTCCAAGTAC-2 | 1 |
1735 rows × 2 columns
Import PKN with CORNETO#
cn.info()
|
|
|
outputs_pkn = list(set(df_edges.parent.tolist()) - set(df_edges.child.tolist()))
inputs_pkn = set(df_edges.child.tolist()) - set(df_edges.parent.tolist())
input_pkn_genes = list(set(g.split("_")[0] for g in inputs_pkn))
len(inputs_pkn), len(outputs_pkn)
(13121, 1)
tuples = [(r.child, 1, r.parent) for _, r in df_edges.iterrows()]
G = cn.Graph.from_sif_tuples(tuples)
G = G.prune(inputs_pkn, outputs_pkn)
# Size of the original PKN provided by the authors
G.shape
(13439, 27579)
Select the single cell data for training#
adata_matched = adata[adata.obs_names.isin(matched_barcodes), adata.var_names.isin(input_pkn_genes)]
adata_matched.shape
(1735, 14229)
non_zero_genes = set(adata_matched.to_df().columns[adata_matched.to_df().sum(axis=0) >= 1e-6].values)
len(non_zero_genes)
12459
len(non_zero_genes.intersection(adata_matched.var_names))
12459
adata_matched = adata_matched[:, adata_matched.var_names.isin(non_zero_genes)]
# Many duplicates still 0 counts
adata_matched = adata_matched[:, adata_matched.to_df().sum(axis=0) != 0]
adata_matched.shape
(1735, 12487)
df_expr = adata_matched.to_df()
df_expr = df_expr.groupby(df_expr.columns, axis=1).max()
df_expr
| A1BG | A2ML1 | AAAS | AACS | AADAT | AAED1 | AAGAB | AAK1 | AAMDC | AAMP | ... | ZSWIM8 | ZUFSP | ZW10 | ZWILCH | ZXDC | ZYG11A | ZYG11B | ZYX | ZZEF1 | ZZZ3 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| AAACCTGCACACATGT-1 | 0.0 | 0.0 | 0.000000 | 0.693147 | 0.0 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 1.098612 | ... | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.0 | 0.000000 | 0.000000 | 0.000000 |
| AAACCTGCACGTCTCT-1 | 0.0 | 0.0 | 0.000000 | 0.000000 | 0.0 | 0.693147 | 0.000000 | 0.000000 | 0.0 | 0.693147 | ... | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.0 | 0.000000 | 0.000000 | 0.000000 |
| AAACCTGTCAATACCG-1 | 0.0 | 0.0 | 0.000000 | 0.000000 | 0.0 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 0.693147 | ... | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.0 | 0.000000 | 0.000000 | 0.000000 |
| AAACCTGTCGTGGTCG-1 | 0.0 | 0.0 | 1.098612 | 0.000000 | 0.0 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 1.098612 | ... | 0.000000 | 0.693147 | 0.000000 | 0.693147 | 0.0 | 0.0 | 0.0 | 0.000000 | 0.000000 | 0.000000 |
| AAACGGGTCTGAGTGT-1 | 0.0 | 0.0 | 0.000000 | 0.000000 | 0.0 | 0.693147 | 0.000000 | 0.000000 | 0.0 | 0.000000 | ... | 0.693147 | 0.000000 | 0.693147 | 0.000000 | 0.0 | 0.0 | 0.0 | 0.000000 | 0.000000 | 0.000000 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| TTTCCTCGTCATGCCG-2 | 0.0 | 0.0 | 0.000000 | 0.000000 | 0.0 | 0.000000 | 0.000000 | 0.693147 | 0.0 | 0.693147 | ... | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.0 | 0.000000 | 0.000000 | 0.000000 |
| TTTGCGCGTAGCCTCG-2 | 0.0 | 0.0 | 1.098612 | 0.000000 | 0.0 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 1.386294 | ... | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.0 | 0.000000 | 0.000000 | 0.000000 |
| TTTGGTTAGATACACA-2 | 0.0 | 0.0 | 1.098612 | 0.000000 | 0.0 | 0.000000 | 0.000000 | 1.098612 | 0.0 | 0.693147 | ... | 0.693147 | 0.000000 | 0.693147 | 0.000000 | 0.0 | 0.0 | 0.0 | 0.693147 | 0.693147 | 0.000000 |
| TTTGGTTGTATGAATG-2 | 0.0 | 0.0 | 0.000000 | 0.000000 | 0.0 | 0.693147 | 1.098612 | 0.000000 | 0.0 | 0.693147 | ... | 0.000000 | 0.693147 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.0 | 0.000000 | 0.000000 | 0.693147 |
| TTTGGTTTCCAAGTAC-2 | 0.0 | 0.0 | 0.000000 | 0.000000 | 0.0 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 0.693147 | ... | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.0 | 1.098612 | 0.000000 | 0.000000 |
1735 rows × 12459 columns
Building and training the KPNN#
Now we will use the provided PKN by the authors and the utility functions in CORNETO to build a KPNN similar to the one used in the original manuscript
import os
os.environ["KERAS_BACKEND"] = "jax"
import keras
from keras.callbacks import EarlyStopping
from sklearn.metrics import (
accuracy_score,
f1_score,
precision_score,
recall_score,
roc_auc_score,
)
from sklearn.model_selection import StratifiedKFold
# Use the data from the experiment
X = df_expr.values
y = df_labels.set_index("barcode").loc[df_expr.index, "TCR"].values
X.shape, y.shape
((1735, 12459), (1735,))
# We can prefilter on top N genes to make this faster
top_n = None
# From the given PKN
outputs_pkn = list(set(df_edges.parent.tolist()) - set(df_edges.child.tolist()))
inputs_pkn = set(df_edges.child.tolist()) - set(df_edges.parent.tolist())
input_pkn_genes = list(set(g.split("_")[0] for g in inputs_pkn))
if top_n is not None and top_n > 0:
input_pkn_genes = list(
set(input_pkn_genes).intersection(df_expr.var(axis=0).sort_values(ascending=False).head(top_n).index)
)
inputs_pkn = list(g + "_gene" for g in input_pkn_genes)
len(inputs_pkn), len(outputs_pkn)
(13121, 1)
input_nn_genes = list(set(input_pkn_genes).intersection(df_expr.columns))
input_nn = [g + "_gene" for g in input_nn_genes]
len(input_nn)
12459
# Build corneto graph
tuples = [(r.child, 1, r.parent) for _, r in df_edges.iterrows()]
G = cn.Graph.from_sif_tuples(tuples)
G = G.prune(input_nn, outputs_pkn)
G.shape
(12767, 25928)
len(input_nn), len(input_nn_genes)
(12459, 12459)
len(set(input_nn).intersection(G.V))
12459
X = df_expr.loc[:, input_nn_genes].values
y = df_labels.set_index("barcode").loc[df_expr.index, "TCR"].values
X.shape, y.shape
((1735, 12459), (1735,))
from corneto._ml import build_dagnn
def stratified_kfold(
G,
inputs,
outputs,
n_splits=5,
shuffle=True,
random_state=42,
lr=0.001,
patience=10,
file_weights="weights",
dagnn_config=dict(
batch_norm_input=True,
batch_norm_center=False,
batch_norm_scale=False,
bias_reg_l1=1e-3,
bias_reg_l2=1e-2,
dropout=0.20,
default_hidden_activation="sigmoid",
default_output_activation="sigmoid",
force_sign=False,
verbose=False,
),
):
kfold = StratifiedKFold(n_splits=n_splits, shuffle=shuffle, random_state=random_state)
models = []
metrics = {m: [] for m in ["accuracy", "precision", "recall", "f1", "roc_auc"]}
for i, (train_idx, val_idx) in enumerate(kfold.split(X, y)):
X_train, X_val = X[train_idx], X[val_idx]
y_train, y_val = y[train_idx], y[val_idx]
print("Building DAG NN model with CORNETO using Keras with JAX...")
print(f" > N. inputs: {len(input_nn)}")
print(f" > N. outputs: {len(outputs_pkn)}")
model = build_dagnn(G, input_nn, outputs_pkn, **dagnn_config)
print(f" > N. parameters: {model.count_params()}")
# Train the model with Adam
opt = keras.optimizers.Adam(learning_rate=lr)
early_stopping = EarlyStopping(monitor="val_loss", patience=patience, restore_best_weights=True)
print("Compiling...")
model.compile(optimizer=opt, loss="binary_crossentropy", metrics=["accuracy"])
print("Fitting...")
model.fit(
X_train,
y_train,
validation_data=(X_val, y_val),
epochs=200,
batch_size=64,
verbose=0,
callbacks=[early_stopping],
)
if file_weights is not None:
filename = f"{file_weights}_{i}.keras"
model.save(filename)
print(f"Weights saved to {filename}")
# Predictions and metrics calculation
y_pred_proba = model.predict(X_val).flatten()
y_pred = (y_pred_proba > 0.5).astype(int)
acc = accuracy_score(y_val, y_pred)
precision = precision_score(y_val, y_pred)
recall = recall_score(y_val, y_pred)
f1 = f1_score(y_val, y_pred)
roc_auc = roc_auc_score(y_val, y_pred_proba)
metrics["accuracy"].append(acc)
metrics["precision"].append(precision)
metrics["recall"].append(recall)
metrics["f1"].append(f1)
metrics["roc_auc"].append(roc_auc)
print(f" > Fold {i} validation ROC-AUC={roc_auc:.3f}")
models.append(model)
return models, metrics
temp_weights = tempfile.mkdtemp()
models, metrics = stratified_kfold(G, input_nn, outputs_pkn, file_weights=os.path.join(temp_weights, "weights"))
print("Validation metrics:")
for k, v in metrics.items():
print(f" - {k}: {np.mean(v):.3f}")
Building DAG NN model with CORNETO using Keras with JAX...
> N. inputs: 12459
> N. outputs: 1
> N. parameters: 26236
Compiling...
Fitting...
Weights saved to /var/folders/b4/gwkwsdb93sv11rtztqbm3l040000gn/T/tmpflwx1756/weights_0.keras
1/11 ━━━━━━━━━━━━━━━━━━━━ 10s 1s/step
10/11 ━━━━━━━━━━━━━━━━━━━━ 0s 18ms/step
11/11 ━━━━━━━━━━━━━━━━━━━━ 0s 113ms/step
11/11 ━━━━━━━━━━━━━━━━━━━━ 2s 113ms/step
> Fold 0 validation ROC-AUC=0.995
Building DAG NN model with CORNETO using Keras with JAX...
> N. inputs: 12459
> N. outputs: 1
> N. parameters: 26236
Compiling...
Fitting...
Weights saved to /var/folders/b4/gwkwsdb93sv11rtztqbm3l040000gn/T/tmpflwx1756/weights_1.keras
1/11 ━━━━━━━━━━━━━━━━━━━━ 10s 1s/step
10/11 ━━━━━━━━━━━━━━━━━━━━ 0s 19ms/step
11/11 ━━━━━━━━━━━━━━━━━━━━ 0s 111ms/step
11/11 ━━━━━━━━━━━━━━━━━━━━ 2s 111ms/step
> Fold 1 validation ROC-AUC=0.978
Building DAG NN model with CORNETO using Keras with JAX...
> N. inputs: 12459
> N. outputs: 1
> N. parameters: 26236
Compiling...
Fitting...
Weights saved to /var/folders/b4/gwkwsdb93sv11rtztqbm3l040000gn/T/tmpflwx1756/weights_2.keras
1/11 ━━━━━━━━━━━━━━━━━━━━ 10s 1s/step
10/11 ━━━━━━━━━━━━━━━━━━━━ 0s 18ms/step
11/11 ━━━━━━━━━━━━━━━━━━━━ 0s 110ms/step
11/11 ━━━━━━━━━━━━━━━━━━━━ 2s 110ms/step
> Fold 2 validation ROC-AUC=0.988
Building DAG NN model with CORNETO using Keras with JAX...
> N. inputs: 12459
> N. outputs: 1
> N. parameters: 26236
Compiling...
Fitting...
Weights saved to /var/folders/b4/gwkwsdb93sv11rtztqbm3l040000gn/T/tmpflwx1756/weights_3.keras
1/11 ━━━━━━━━━━━━━━━━━━━━ 10s 1s/step
10/11 ━━━━━━━━━━━━━━━━━━━━ 0s 20ms/step
11/11 ━━━━━━━━━━━━━━━━━━━━ 0s 112ms/step
11/11 ━━━━━━━━━━━━━━━━━━━━ 2s 112ms/step
> Fold 3 validation ROC-AUC=0.993
Building DAG NN model with CORNETO using Keras with JAX...
> N. inputs: 12459
> N. outputs: 1
> N. parameters: 26236
Compiling...
Fitting...
Weights saved to /var/folders/b4/gwkwsdb93sv11rtztqbm3l040000gn/T/tmpflwx1756/weights_4.keras
1/11 ━━━━━━━━━━━━━━━━━━━━ 11s 1s/step
10/11 ━━━━━━━━━━━━━━━━━━━━ 0s 19ms/step
11/11 ━━━━━━━━━━━━━━━━━━━━ 0s 112ms/step
11/11 ━━━━━━━━━━━━━━━━━━━━ 2s 113ms/step
> Fold 4 validation ROC-AUC=0.993
Validation metrics:
- accuracy: 0.961
- precision: 0.967
- recall: 0.954
- f1: 0.960
- roc_auc: 0.989
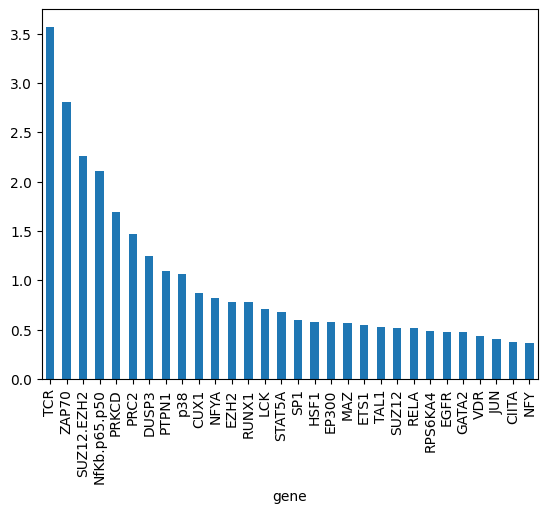
Now, we will analyze the learned biases for each of the nodes in the graph. Note that authors in the KPNN paper explain a way to extract weights for the nodes, based on the learned interactions and accounting for biases in the structure of the NN. Here we just show the learned biases of the nodes of the NN across 5 folds. Please be careful interpreting these weights.
# We collect the weights obtained in each fold
def load_biases(file="weights", folds=5):
biases = []
mean_inputs = []
for i in range(5):
model = keras.models.load_model(f"{file}_{i}.keras")
for layer in model.layers:
weights = layer.get_weights()
if weights:
biases.append((i, layer.name, weights[1][0]))
mean_inputs.append((i, layer.name, weights[0].mean()))
df_biases = pd.DataFrame(biases, columns=["fold", "gene", "bias"])
df_biases["abs_bias"] = df_biases.bias.abs()
df_biases["pow2_bias"] = df_biases.bias.pow(2)
df_biases = df_biases.set_index(["fold", "gene"])
return df_biases
df_biases = load_biases(file=os.path.join(temp_weights, "weights"), folds=5)
df_biases.sort_values(by="abs_bias", ascending=False).head(10)
| bias | abs_bias | pow2_bias | ||
|---|---|---|---|---|
| fold | gene | |||
| 2 | SUZ12.EZH2 | -2.319227 | 2.319227 | 5.378813 |
| 3 | TCR | -2.314526 | 2.314526 | 5.357029 |
| 1 | ZAP70 | -2.168838 | 2.168838 | 4.703859 |
| 2 | TCR | 2.163686 | 2.163686 | 4.681535 |
| 3 | NfKb.p65.p50 | -2.100779 | 2.100779 | 4.413271 |
| 4 | SUZ12.EZH2 | -1.978842 | 1.978842 | 3.915815 |
| 3 | ZAP70 | 1.965354 | 1.965354 | 3.862615 |
| 4 | PRC2 | -1.936076 | 1.936076 | 3.748389 |
| 0 | TCR | 1.794367 | 1.794367 | 3.219754 |
| 1 | NfKb.p65.p50 | -1.788792 | 1.788792 | 3.199777 |
df_biases_full = df_biases.copy().reset_index()
gene_biases_score = df_biases_full.groupby("gene")["pow2_bias"].mean().sort_values(ascending=False)
gene_biases_score
gene
TCR 3.567696
ZAP70 2.803867
SUZ12.EZH2 2.255738
NfKb.p65.p50 2.109870
PRKCD 1.688119
...
CBX3 0.000156
KMT2D 0.000054
HMT 0.000016
GATA3 0.000015
SRY 0.000011
Name: pow2_bias, Length: 308, dtype: float32
gene_biases_score.head(30).plot.bar()
<Axes: xlabel='gene'>
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
# Get the top genes sorted by mean bias
top_genes = gene_biases_score.head(30).index
# Filter the DataFrame to include only rows with these top genes
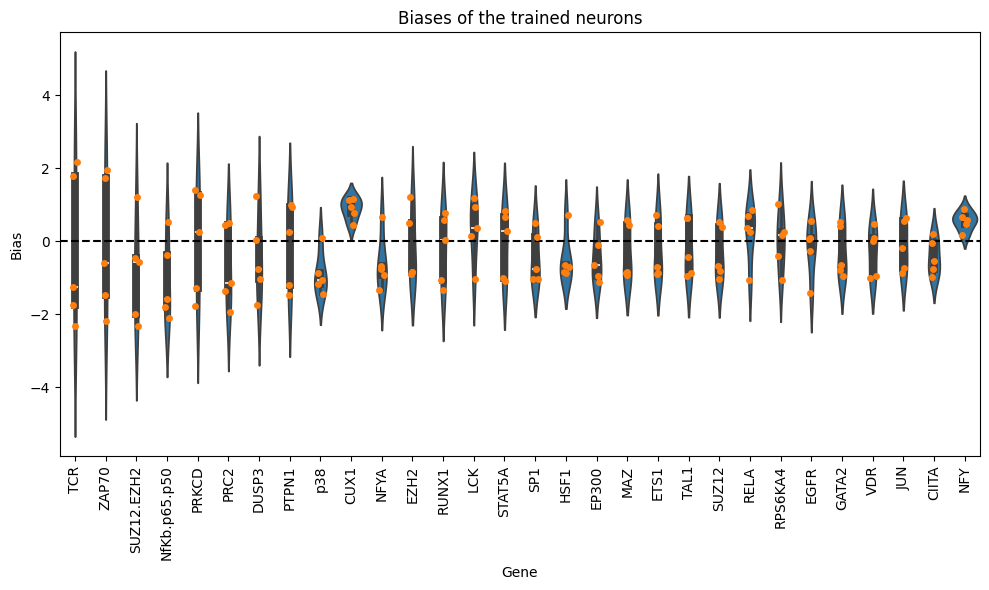
filtered_df = df_biases_full[df_biases_full["gene"].isin(top_genes)]
plt.figure(figsize=(10, 6))
sns.violinplot(data=filtered_df, x="gene", y="bias", order=top_genes, dodge=True)
sns.stripplot(data=filtered_df, x="gene", y="bias", order=top_genes, dodge=True)
plt.xticks(rotation=90)
plt.title("Biases of the trained neurons")
plt.xlabel("Gene")
plt.ylabel("Bias")
plt.axhline(0, linestyle="--", color="k")
plt.tight_layout()
Use CORNETO for NN pruning#
Now we will show how CORNETO can be used to extract a smaller, yet complete DAG from the original PKN provided by the authors. We will add input edges to each input node and an output edge through TCR to indicate which nodes are the inputs and which one the output. We will use then Acyclic Flow to find the smallest DAG comprising these nodes
G_dag = G.copy()
new_edges = []
for g in input_nn:
new_edges.append(G_dag.add_edge((), g))
new_edges.append(G_dag.add_edge("TCR", ()))
print(G_dag.shape)
# Find small DAG. We use Acyclic Flow to find over the space of DAGs
P = cn.opt.AcyclicFlow(G_dag)
# We enforce that the input genes and the output gene are part of the solution
P += P.expr.with_flow[new_edges] == 1
# Minimize the number of active edges
P.add_objectives(sum(P.expr.with_flow), weights=1)
P.solve(solver="HIGHS", verbosity=1, max_seconds=300);
(12767, 38388)
===============================================================================
CVXPY
v1.7.5
===============================================================================
-------------------------------------------------------------------------------
Compilation
-------------------------------------------------------------------------------
-------------------------------------------------------------------------------
Numerical solver
-------------------------------------------------------------------------------
Running HiGHS 1.13.0 (git hash: 1bce6d5): Copyright (c) 2026 under MIT licence terms
MIP has 332945 rows; 127931 cols; 628290 nonzeros; 76776 integer variables (76776 binary)
Coefficient ranges:
Matrix [1e-04, 1e+04]
Cost [1e+00, 1e+00]
Bound [1e+00, 1e+00]
RHS [1e+00, 1e+04]
Presolving model
61423 rows, 50344 cols, 162081 nonzeros 0s
41020 rows, 42186 cols, 129115 nonzeros 0s
39333 rows, 38152 cols, 131120 nonzeros 0s
Presolve reductions: rows 39333(-293612); columns 38152(-89779); nonzeros 131120(-497170)
Objective function is integral with scale 1
Solving MIP model with:
39333 rows
38152 cols (18820 binary, 0 integer, 0 implied int., 19332 continuous, 0 domain fixed)
131120 nonzeros
Src: B => Branching; C => Central rounding; F => Feasibility pump; H => Heuristic;
I => Shifting; J => Feasibility jump; L => Sub-MIP; P => Empty MIP; R => Randomized rounding;
S => Solve LP; T => Evaluate node; U => Unbounded; X => User solution; Y => HiGHS solution;
Z => ZI Round; l => Trivial lower; p => Trivial point; u => Trivial upper; z => Trivial zero
Nodes | B&B Tree | Objective Bounds | Dynamic Constraints | Work
Src Proc. InQueue | Leaves Expl. | BestBound BestSol Gap | Cuts InLp Confl. | LpIters Time
0 0 0 0.00% 19719 inf inf 0 0 0 0 0.9s
0 0 0 0.00% 19719 inf inf 0 0 5 16346 1.8s
0 0 0 0.00% 24886.380366 inf inf 6492 5461 58 75101 22.7s
L 0 0 0 0.00% 24924.373044 25087 0.65% 8224 5729 113 77722 36.5s
0.2% inactive integer columns, restarting
Model after restart has 39257 rows, 38106 cols (18777 bin., 0 int., 0 impl., 19329 cont., 0 dom.fix.), and 130968 nonzeros
0 0 0 0.00% 24924.373044 25087 0.65% 5499 0 0 142175 36.9s
0 0 0 0.00% 24924.373059 25087 0.65% 5499 5433 4 254650 69.2s
Symmetry detection completed in 4.6s
Found 2475 generator(s)
48 0 1 0.00% 24927.248967 25087 0.64% 8027 5631 99 322948 100.7s
192 188 1 0.00% 24996.054495 25087 0.36% 8054 5631 170 337853 113.6s
232 187 2 0.00% 24996.054495 25087 0.36% 8482 5712 180 410758 151.7s
370 309 6 0.00% 24996.054495 25087 0.36% 8831 5745 211 429962 162.5s
T 401 239 7 0.00% 24996.054495 25079 0.33% 8831 5745 216 430098 166.5s
503 359 9 0.00% 24996.054495 25079 0.33% 9205 5774 225 451338 176.4s
T 503 311 9 0.00% 24996.054495 25073 0.31% 9205 5774 229 451338 178.6s
L 577 331 10 0.00% 24996.054495 25067 0.28% 9409 5474 231 453184 185.6s
630 331 11 0.00% 24996.054495 25067 0.28% 9411 5474 239 479529 192.5s
766 430 13 0.00% 24997.054475 25067 0.28% 9497 5488 261 501683 205.1s
853 595 13 0.00% 24997.054485 25067 0.28% 9676 5515 283 529168 220.8s
905 594 14 0.00% 24997.054485 25067 0.28% 10056 5493 293 561558 236.5s
L 935 632 14 0.00% 24997.054485 25066 0.28% 10295 5510 299 563727 244.6s
984 632 15 0.00% 24997.054485 25066 0.28% 10295 5510 308 589295 249.6s
Nodes | B&B Tree | Objective Bounds | Dynamic Constraints | Work
Src Proc. InQueue | Leaves Expl. | BestBound BestSol Gap | Cuts InLp Confl. | LpIters Time
1068 692 16 0.00% 24997.054485 25066 0.28% 10386 5524 323 649063 279.6s
1139 776 17 0.00% 24997.054485 25066 0.28% 10392 5538 336 685024 299.3s
1151 838 18 0.00% 25006.0518 25066 0.24% 10392 5538 341 687194 300.0s
Solving report
Status Time limit reached
Primal bound 25066
Dual bound 25007
Gap 0.235% (tolerance: 0.01%)
P-D integral 1.06568195332
Solution status feasible
25066 (objective)
0 (bound viol.)
0 (int. viol.)
0 (row viol.)
Timing 300.02
Max sub-MIP depth 10
Nodes 1151
Repair LPs 0
LP iterations 687194
146288 (strong br.)
95062 (separation)
193529 (heuristics)
-------------------------------------------------------------------------------
Summary
-------------------------------------------------------------------------------
G_subdag = G_dag.edge_subgraph(P.expr.with_flow.value > 0.5)
G_dag.shape, G_subdag.shape
((12767, 38388), (12607, 25066))
rel_dag_compression = (1 - (G_subdag.num_edges / G_dag.num_edges)) * 100
print(f"KPNN edge compression (0-100%): {rel_dag_compression:.2f}%")
KPNN edge compression (0-100%): 34.70%
pruned_models, pruned_metrics = stratified_kfold(
G_subdag,
input_nn,
outputs_pkn,
file_weights=os.path.join(temp_weights, "pruned_weights"),
)
print("Validation metrics:")
for k, v in pruned_metrics.items():
print(f" - {k}: {np.mean(v):.3f}")
Building DAG NN model with CORNETO using Keras with JAX...
> N. inputs: 12459
> N. outputs: 1
> N. parameters: 12754
Compiling...
Fitting...
Weights saved to /var/folders/b4/gwkwsdb93sv11rtztqbm3l040000gn/T/tmpflwx1756/pruned_weights_0.keras
1/11 ━━━━━━━━━━━━━━━━━━━━ 5s 539ms/step
10/11 ━━━━━━━━━━━━━━━━━━━━ 0s 10ms/step
11/11 ━━━━━━━━━━━━━━━━━━━━ 0s 53ms/step
11/11 ━━━━━━━━━━━━━━━━━━━━ 1s 53ms/step
> Fold 0 validation ROC-AUC=0.988
Building DAG NN model with CORNETO using Keras with JAX...
> N. inputs: 12459
> N. outputs: 1
> N. parameters: 12754
Compiling...
Fitting...
Weights saved to /var/folders/b4/gwkwsdb93sv11rtztqbm3l040000gn/T/tmpflwx1756/pruned_weights_1.keras
1/11 ━━━━━━━━━━━━━━━━━━━━ 5s 525ms/step
10/11 ━━━━━━━━━━━━━━━━━━━━ 0s 10ms/step
11/11 ━━━━━━━━━━━━━━━━━━━━ 0s 53ms/step
11/11 ━━━━━━━━━━━━━━━━━━━━ 1s 53ms/step
> Fold 1 validation ROC-AUC=0.980
Building DAG NN model with CORNETO using Keras with JAX...
> N. inputs: 12459
> N. outputs: 1
> N. parameters: 12754
Compiling...
Fitting...
Weights saved to /var/folders/b4/gwkwsdb93sv11rtztqbm3l040000gn/T/tmpflwx1756/pruned_weights_2.keras
1/11 ━━━━━━━━━━━━━━━━━━━━ 5s 544ms/step
10/11 ━━━━━━━━━━━━━━━━━━━━ 0s 11ms/step
11/11 ━━━━━━━━━━━━━━━━━━━━ 0s 53ms/step
11/11 ━━━━━━━━━━━━━━━━━━━━ 1s 53ms/step
> Fold 2 validation ROC-AUC=0.987
Building DAG NN model with CORNETO using Keras with JAX...
> N. inputs: 12459
> N. outputs: 1
> N. parameters: 12754
Compiling...
Fitting...
Weights saved to /var/folders/b4/gwkwsdb93sv11rtztqbm3l040000gn/T/tmpflwx1756/pruned_weights_3.keras
1/11 ━━━━━━━━━━━━━━━━━━━━ 5s 513ms/step
10/11 ━━━━━━━━━━━━━━━━━━━━ 0s 11ms/step
11/11 ━━━━━━━━━━━━━━━━━━━━ 0s 53ms/step
11/11 ━━━━━━━━━━━━━━━━━━━━ 1s 53ms/step
> Fold 3 validation ROC-AUC=0.992
Building DAG NN model with CORNETO using Keras with JAX...
> N. inputs: 12459
> N. outputs: 1
> N. parameters: 12754
Compiling...
Fitting...
Weights saved to /var/folders/b4/gwkwsdb93sv11rtztqbm3l040000gn/T/tmpflwx1756/pruned_weights_4.keras
1/11 ━━━━━━━━━━━━━━━━━━━━ 5s 526ms/step
10/11 ━━━━━━━━━━━━━━━━━━━━ 0s 10ms/step
11/11 ━━━━━━━━━━━━━━━━━━━━ 0s 54ms/step
11/11 ━━━━━━━━━━━━━━━━━━━━ 1s 54ms/step
> Fold 4 validation ROC-AUC=0.994
Validation metrics:
- accuracy: 0.963
- precision: 0.969
- recall: 0.954
- f1: 0.962
- roc_auc: 0.988
df_biases_pruned = load_biases(file=os.path.join(temp_weights, "pruned_weights"), folds=5)
df_biases_pruned.sort_values(by="abs_bias", ascending=False).head(10)
| bias | abs_bias | pow2_bias | ||
|---|---|---|---|---|
| fold | gene | |||
| 2 | ZAP70 | -2.833718 | 2.833718 | 8.029957 |
| 3 | SUZ12.EZH2 | -2.608386 | 2.608386 | 6.803678 |
| 4 | PRKCD | -2.220839 | 2.220839 | 4.932124 |
| 3 | TCR | 2.169856 | 2.169856 | 4.708274 |
| 4 | TCR | -2.168031 | 2.168031 | 4.700360 |
| ZAP70 | 2.140123 | 2.140123 | 4.580125 | |
| 3 | ZAP70 | 2.107303 | 2.107303 | 4.440725 |
| 2 | NFYA | 2.093171 | 2.093171 | 4.381363 |
| 1 | NFYA | 2.036027 | 2.036027 | 4.145407 |
| 2 | MAX | -2.015005 | 2.015005 | 4.060244 |
df_biases_prunedr = df_biases_pruned.copy().reset_index()
gene_biases_score = df_biases_prunedr.groupby("gene")["pow2_bias"].mean().sort_values(ascending=False)
gene_biases_score
gene
ZAP70 4.356112
TCR 3.455697
NFYA 3.271365
SUZ12.EZH2 2.938754
PRKCD 2.340390
...
ZNF217 0.003361
MLL.SET.subcomplex 0.002986
REST 0.000634
ASH2L 0.000447
GATA3 0.000002
Name: pow2_bias, Length: 148, dtype: float32
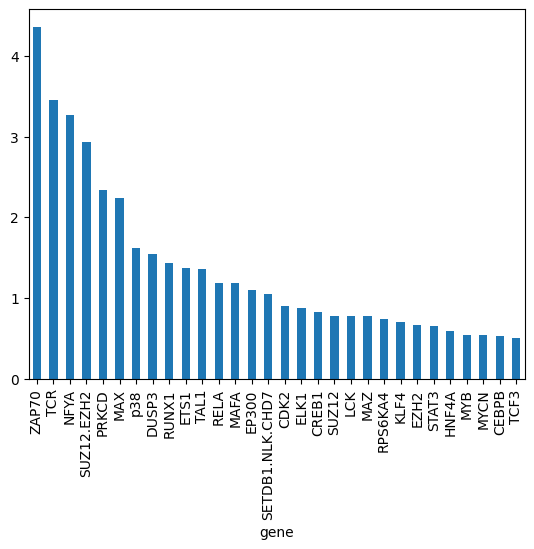
gene_biases_score.head(30).plot.bar()
<Axes: xlabel='gene'>
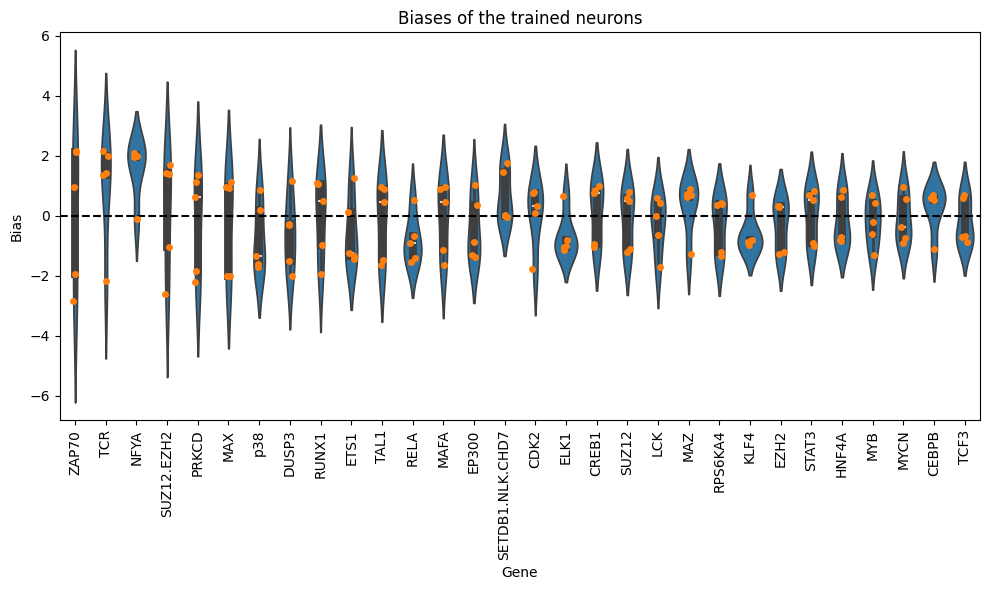
# Get the top genes sorted by mean bias
top_genes = gene_biases_score.head(30).index
# Filter the DataFrame to include only rows with these top genes
filtered_df = df_biases_prunedr[df_biases_prunedr["gene"].isin(top_genes)]
plt.figure(figsize=(10, 6))
sns.violinplot(data=filtered_df, x="gene", y="bias", order=top_genes, dodge=True)
sns.stripplot(data=filtered_df, x="gene", y="bias", order=top_genes, dodge=True)
plt.xticks(rotation=90)
plt.title("Biases of the trained neurons")
plt.xlabel("Gene")
plt.ylabel("Bias")
plt.axhline(0, linestyle="--", color="k")
plt.tight_layout()
param_compression = (1 - (pruned_models[0].count_params() / models[0].count_params())) * 100
print(f"Parameter compression: {param_compression:.2f}%")
Parameter compression: 51.39%
perf_degradation = (
(np.mean(metrics["roc_auc"]) - np.mean(pruned_metrics["roc_auc"])) / np.mean(metrics["roc_auc"])
) * 100
print(
f"Degradation in ROC-AUC after compression (positive = decrease in performance, negative = increase in performance): {perf_degradation:.2f}%"
)
Degradation in ROC-AUC after compression (positive = decrease in performance, negative = increase in performance): 0.11%
