Multi-sample PCST#
Here we demonstrate the use of the multi-sample PCST method, and we also compare against the standard single sample PCST approximation implemented in the pcst_fast package
import random
import matplotlib.pyplot as plt
import networkx as nx
import numpy as np
import pcst_fast # Ensure pcst_fast is installed
def generate_random_graph(num_nodes=100, probability=0.1, num_terminals=10):
"""Generates a random unweighted graph using the Erdős-Rényi model and assigns terminal nodes.
Prizes are not assigned during generation.
Args:
num_nodes (int): Number of nodes in the graph.
probability (float): Probability for edge creation.
num_terminals (int): Number of nodes to mark as terminals.
Returns:
G (networkx.Graph): Graph with a 'terminal' attribute on nodes.
"""
G = nx.erdos_renyi_graph(num_nodes, probability)
# Randomly select terminal nodes
terminals = random.sample(list(G.nodes()), min(num_terminals, G.number_of_nodes()))
for node in G.nodes():
G.nodes[node]["terminal"] = node in terminals
return G
def assign_random_costs_and_prizes(G, num_prized=10, prize_range=(1, 50)):
"""Creates a new copy of the given graph, assigns random edge costs and prizes.
The original graph remains unchanged. Edge costs are stored in the 'value' attribute.
Only a specified number of non-terminal nodes receive a random prize; all other nodes are assigned a prize of 0.
Args:
G (networkx.Graph): The input graph whose topology is preserved.
num_prized (int): The number of non-terminal nodes to receive a prize.
prize_range (tuple): Tuple (min, max) for random prize values.
Returns:
G_new (networkx.Graph): A new graph with updated 'value' attributes on edges and 'prize' attributes on nodes.
"""
# Create a copy of the graph so the original is not modified.
G_new = G.copy()
# Assign random edge costs
for u, v in G_new.edges():
G_new[u][v]["value"] = random.uniform(1, 10)
# Identify non-terminal nodes
non_terminal_nodes = [node for node in G_new.nodes() if not G_new.nodes[node].get("terminal", False)]
# Randomly select nodes to assign prizes (do not exceed available nodes)
prized_nodes = random.sample(non_terminal_nodes, min(num_prized, len(non_terminal_nodes)))
# Assign prizes: nodes selected get a random prize, all others get 0
for node in G_new.nodes():
if node in prized_nodes:
G_new.nodes[node]["prize"] = random.uniform(*prize_range)
else:
G_new.nodes[node]["prize"] = 0
return G_new
def solve_pcst_with_pcst_fast(G, root=-1, num_clusters=1, pruning="strong", verbosity=0):
"""Solves the PCST problem on a NetworkX graph using the pcst_fast package.
Args:
G (networkx.Graph): Graph with 'value' on edges and 'prize' on nodes.
root (int): Index of the root node (or -1 for unrooted).
num_clusters (int): Desired number of clusters in the solution.
pruning (str): Pruning method ('none', 'simple', 'gw', 'strong').
verbosity (int): Verbosity level.
Returns:
selected_nodes (list): Node IDs included in the PCST solution.
selected_edges (list): Edge tuples included in the solution.
"""
node_list = list(G.nodes())
node_index_map = {node: i for i, node in enumerate(node_list)}
index_node_map = {i: node for node, i in node_index_map.items()}
edge_list = []
edge_costs = []
for u, v, data in G.edges(data=True):
edge_list.append([node_index_map[u], node_index_map[v]])
edge_costs.append(data["value"])
prizes = np.array([G.nodes[node].get("prize", 0) for node in node_list], dtype=np.float64)
edges_array = np.array(edge_list, dtype=np.int64)
costs_array = np.array(edge_costs, dtype=np.float64)
vertices_out, edges_out = pcst_fast.pcst_fast(
edges_array, prizes, costs_array, root, num_clusters, pruning, verbosity
)
selected_nodes = [index_node_map[i] for i in vertices_out]
selected_edges = [tuple(node_list[idx] for idx in edge_list[i]) for i in edges_out]
return selected_nodes, selected_edges
def compute_pcst_cost(G, selected_edges):
"""Calculates the cost components and total PCST objective value based solely on the selected edges.
The function:
- Derives the set of selected nodes from the provided edges.
- Builds a graph using only the selected edges.
- Checks that the selected edges form a single connected subgraph.
Returns:
dict with keys 'edge_cost', 'prize_collected', and 'objective'
Raises:
ValueError: If any provided edge is not in G or if the selected edges do not form a connected subgraph.
"""
# Derive selected nodes from edges and verify edge validity
selected_nodes = set()
for u, v in selected_edges:
if not G.has_edge(u, v):
raise ValueError(f"Edge ({u}, {v}) is not present in the graph.")
selected_nodes.update([u, v])
# For non-empty selections, check connectivity using only the selected edges.
if selected_edges:
H = nx.Graph()
H.add_nodes_from(selected_nodes)
H.add_edges_from(selected_edges)
if not nx.is_connected(H):
raise ValueError("Selected edges do not form a connected subgraph.")
# Calculate the total edge cost.
edge_cost = sum(G[u][v]["value"] for u, v in selected_edges)
# Sum the prizes for all nodes in the connected subgraph.
prize_collected = sum(G.nodes[n].get("prize", 0) for n in selected_nodes)
objective = edge_cost - prize_collected
return {
"edge_cost": edge_cost,
"prize_collected": prize_collected,
"objective": objective,
}
def visualize(G, selected_nodes, selected_edges):
pos = nx.forceatlas2_layout(G, seed=42)
plt.figure(figsize=(8, 8))
# Base layer: faint background graph
nx.draw_networkx_edges(G, pos, alpha=0.5, width=0.5, edge_color="lightgray")
nx.draw_networkx_nodes(G, pos, node_size=20, node_color="lightgray", alpha=0.3)
# Selected edges (solution)
nx.draw_networkx_edges(G, pos, edgelist=selected_edges, width=2.5, edge_color="#007acc", alpha=0.9)
# Selected nodes (solution)
nx.draw_networkx_nodes(
G,
pos,
nodelist=selected_nodes,
node_size=150,
node_color="#d62728",
edgecolors="white",
linewidths=0.8,
)
# Optional: add labels or prizes
node_labels = {n: f"{(G.nodes[n].get('prize', 0)):.2f}" for n in selected_nodes}
nx.draw_networkx_labels(G, pos, labels=node_labels, font_size=8, font_color="black")
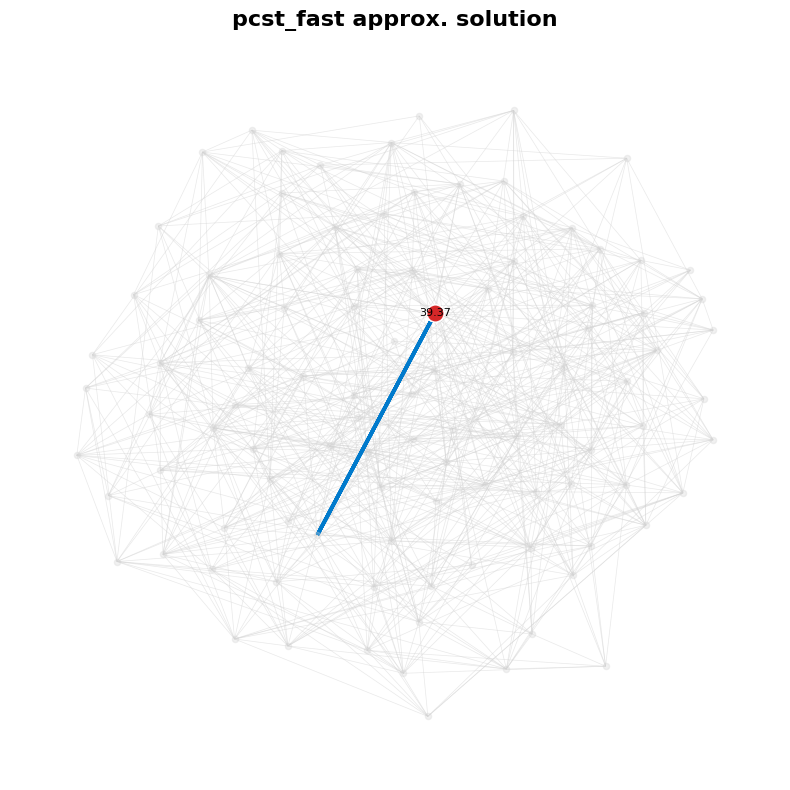
plt.title("pcst_fast approx. solution", fontsize=16, fontweight="bold", pad=15)
plt.axis("off")
plt.tight_layout()
plt.show()
random.seed(10)
# 1. Generate the base graph with a fixed topology (no prizes assigned).
G_original = generate_random_graph(num_nodes=100, probability=0.15, num_terminals=5)
# 2. Create two different samples
G_0 = assign_random_costs_and_prizes(G_original, num_prized=8, prize_range=(1, 50))
G_1 = assign_random_costs_and_prizes(G_original, num_prized=12, prize_range=(1, 50))
# 3. Get an heuristic solution with pcst_fast
selected_nodes_0, selected_edges_0 = solve_pcst_with_pcst_fast(G_0)
selected_nodes_1, selected_edges_1 = solve_pcst_with_pcst_fast(G_1)
compute_pcst_cost(G_0, selected_edges_0), compute_pcst_cost(G_1, selected_edges_1)
({'edge_cost': 15.234903706826126,
'prize_collected': 88.96918230765598,
'objective': -73.73427860082985},
{'edge_cost': 17.372038611342052,
'prize_collected': 47.12252810601924,
'objective': -29.750489494677186})
visualize(G_0, selected_nodes_0, selected_edges_0)
Using CORNETO#
import corneto as cn
cn.info()
|
|
|
from corneto.utils import check_gurobi
check_gurobi(raise_on_error=True)
---------------------------------------------------------------------------
ModuleNotFoundError Traceback (most recent call last)
File ~/work/corneto/corneto/corneto/utils/_solvers.py:8, in check_gurobi(verbose, solver_verbose, raise_on_error)
7 try:
----> 8 import gurobipy as gp
9 except ImportError:
ModuleNotFoundError: No module named 'gurobipy'
During handling of the above exception, another exception occurred:
ImportError Traceback (most recent call last)
Cell In[4], line 3
1 from corneto.utils import check_gurobi
----> 3 check_gurobi(raise_on_error=True)
File ~/work/corneto/corneto/corneto/utils/_solvers.py:10, in check_gurobi(verbose, solver_verbose, raise_on_error)
8 import gurobipy as gp
9 except ImportError:
---> 10 raise ImportError("Gurobipy is not installed. Please install Gurobi and its Python bindings.")
11 if verbose:
12 print("Gurobipy successfully imported.")
ImportError: Gurobipy is not installed. Please install Gurobi and its Python bindings.
from corneto.contrib.networkx import networkx_to_corneto_graph
Gc0 = networkx_to_corneto_graph(G_0)
Gc0.shape
(100, 685)
from corneto._data import Data
def get_data(G):
features = []
for i, attr in enumerate(G.get_attr_edges()):
features.append(dict(id=i, mapping="edge", value=attr.value))
for v in G.V:
attr = G.get_attr_vertex(v)
if attr["terminal"] and not attr.get("prize", None):
# features.append(dict(id=v, mapping="vertex", role="terminal"))
pass
elif attr.get("prize", None):
features.append(dict(id=v, mapping="vertex", role="prize", value=attr["prize"]))
D = Data.from_dict({"s1": {"features": features}})
return D
def get_edge_list(graph):
edges = []
for s, t in graph.E:
s = list(s)
t = list(t)
if len(s) > 0:
s = list(s)[0]
else:
continue
if len(t) > 0:
t = list(t)[0]
else:
continue
edges.append((s, t))
return edges
D = get_data(Gc0)
D
Data(n_samples=1, n_feats=[693])
from corneto.methods.future.pcst import PrizeCollectingSteinerTree as PCST
m = PCST(lambda_reg=0, strict_acyclic=True)
P = m.build(Gc0, D)
P.solve(solver="gurobi", verbosity=0, TimeLimit=300);
Set parameter Username
Set parameter LicenseID to value 2593994
Academic license - for non-commercial use only - expires 2025-12-02
for o in P.objectives:
print(o.value)
25.822958394858087
235.26839405807772
18.0
P.objectives[0].value - P.objectives[1].value
-209.44543566321963
G0_sol = m.processed_graph.edge_subgraph(np.flatnonzero(P.expr.with_flow.value > 0.5))
compute_pcst_cost(G_0, get_edge_list(G0_sol))
{'edge_cost': 25.822958394858087,
'prize_collected': 235.26839405807772,
'objective': -209.44543566321963}
Gc1 = networkx_to_corneto_graph(G_1)
m = PCST(lambda_reg=0, strict_acyclic=True)
P = m.build(Gc1, get_data(Gc1))
P.solve(solver="gurobi", verbosity=0, TimeLimit=300);
G1_sol = m.processed_graph.edge_subgraph(np.flatnonzero(P.expr.with_flow.value > 0.5))
compute_pcst_cost(G_1, get_edge_list(G1_sol))
{'edge_cost': 36.85287997763584,
'prize_collected': 315.445743958666,
'objective': -278.5928639810302}
# pcst_fast sol
compute_pcst_cost(G_1, selected_edges_1)
{'edge_cost': 34.323495165255245,
'prize_collected': 310.25687273661805,
'objective': -275.9333775713628}
Multi-sample PCST#
Now we use multi-sample PCST to simultaneously solve both problems, using the structured sparsity regularization.
# Lets combine both problems into one multi PCST
dataset = get_data(Gc0)
dataset.add_sample("s2", get_data(Gc1).samples["s1"])
dataset
Data(n_samples=2, n_feats=[693 697])
Gc_original = networkx_to_corneto_graph(G_original)
m = PCST(lambda_reg=0, strict_acyclic=True, root_selection_strategy="best")
P = m.build(Gc_original, dataset)
P.solve(solver="gurobi", verbosity=0, TimeLimit=30);
# With no regularization, solutions should be the same
G0_sol = m.processed_graph.edge_subgraph(np.flatnonzero(P.expr.with_flow.value[:, 0] > 0.5))
compute_pcst_cost(G_0, get_edge_list(G0_sol))
{'edge_cost': 25.822958394858087,
'prize_collected': 235.26839405807772,
'objective': -209.44543566321963}
G1_sol = m.processed_graph.edge_subgraph(np.flatnonzero(P.expr.with_flow.value[:, 1] > 0.5))
compute_pcst_cost(G_1, get_edge_list(G1_sol))
{'edge_cost': 36.85287997763584,
'prize_collected': 315.445743958666,
'objective': -278.5928639810302}
m.processed_graph.edge_subgraph(np.flatnonzero(P.expr.with_flow.value[:, 0] > 0.5)).plot(layout="neato")
m.processed_graph.edge_subgraph(np.flatnonzero(P.expr.with_flow.value[:, 1] > 0.5)).plot(layout="neato")
Testing the effect of \(\lambda\)#
We now evaluate the impact of the structured sparsity-inducing penalty, which encourages entire sets of edges to be excluded across all samples. This penalty helps enforce group-level sparsity, effectively removing certain features from all models simultaneously. To speed up the optimization process, we also leverage Gurobi’s warm-start capability. By setting warm_start = True, we can adjust the penalty parameter \(\lambda\) and resolve the problem more efficiently, without needing to rebuild the entire optimization model from scratch.
def jaccard_similarity(a, b):
intersection = np.logical_and(a, b).sum()
union = np.logical_or(a, b).sum()
if union == 0:
return 1.0 # Define similarity as 1 if both are all zeros
return intersection / union
def hamming_similarity(a, b):
return (a == b).sum() / len(a)
results = []
for l in [
0.0,
0.5,
1.0,
1.5,
2.0,
2.5,
3.0,
3.5,
4.0,
5.0,
10.0,
15.0,
20.0,
25.0,
30.0,
]:
print(f"Lambda = {l}")
m.lambda_reg_param.value = l
P.solve(solver="gurobi", warm_start=True, verbosity=0, TimeLimit=120)
sol0 = m.processed_graph.edge_subgraph(np.flatnonzero(P.expr.with_flow.value[:, 0] > 0.5))
sol1 = m.processed_graph.edge_subgraph(np.flatnonzero(P.expr.with_flow.value[:, 1] > 0.5))
# Network union size
size_0 = np.sum(P.expr.with_flow.value[:, 0] > 0.5)
size_1 = np.sum(P.expr.with_flow.value[:, 1] > 0.5)
union_size = np.sum(P.expr.with_flow.value > 0.5, axis=1).sum()
score1 = compute_pcst_cost(G_0, get_edge_list(sol0))
score2 = compute_pcst_cost(G_1, get_edge_list(sol1))
jaccard = jaccard_similarity(P.expr.with_flow.value[:, 0], P.expr.with_flow.value[:, 1])
hamming = hamming_similarity(P.expr.with_flow.value[:, 0], P.expr.with_flow.value[:, 1])
results.append(
(
l,
score1["objective"],
score2["objective"],
jaccard,
hamming,
union_size,
size_0,
size_1,
)
)
print(" > Obj. sample 1:", score1)
print(" > Obj. sample 2:", score2)
print(" > Jaccard dist.:", jaccard)
print(" > Hamming dist.:", hamming)
Lambda = 0.0
> Obj. sample 1: {'edge_cost': 25.822958394858087, 'prize_collected': 235.26839405807772, 'objective': -209.44543566321963}
> Obj. sample 2: {'edge_cost': 36.85287997763584, 'prize_collected': 315.445743958666, 'objective': -278.5928639810302}
> Jaccard dist.: 0.02127659574468085
> Hamming dist.: 0.9346590909090909
Lambda = 0.5
> Obj. sample 1: {'edge_cost': 25.822958394858087, 'prize_collected': 235.26839405807772, 'objective': -209.44543566321963}
> Obj. sample 2: {'edge_cost': 32.57672368032401, 'prize_collected': 310.25687273661805, 'objective': -277.68014905629406}
> Jaccard dist.: 0.022727272727272728
> Hamming dist.: 0.9389204545454546
Lambda = 1.0
> Obj. sample 1: {'edge_cost': 25.822958394858087, 'prize_collected': 235.26839405807772, 'objective': -209.44543566321963}
> Obj. sample 2: {'edge_cost': 33.23852158520403, 'prize_collected': 310.25687273661805, 'objective': -277.01835115141404}
> Jaccard dist.: 0.023255813953488372
> Hamming dist.: 0.9403409090909091
Lambda = 1.5
> Obj. sample 1: {'edge_cost': 25.822958394858087, 'prize_collected': 235.26839405807772, 'objective': -209.44543566321963}
> Obj. sample 2: {'edge_cost': 32.675953291132046, 'prize_collected': 306.3730702170471, 'objective': -273.69711692591505}
> Jaccard dist.: 0.025
> Hamming dist.: 0.9446022727272727
Lambda = 2.0
> Obj. sample 1: {'edge_cost': 27.354768134907065, 'prize_collected': 235.26839405807772, 'objective': -207.91362592317066}
> Obj. sample 2: {'edge_cost': 32.675953291132046, 'prize_collected': 306.3730702170471, 'objective': -273.69711692591505}
> Jaccard dist.: 0.02564102564102564
> Hamming dist.: 0.9460227272727273
Lambda = 2.5
> Obj. sample 1: {'edge_cost': 27.354768134907065, 'prize_collected': 235.26839405807772, 'objective': -207.91362592317066}
> Obj. sample 2: {'edge_cost': 32.675953291132046, 'prize_collected': 306.3730702170471, 'objective': -273.69711692591505}
> Jaccard dist.: 0.02564102564102564
> Hamming dist.: 0.9460227272727273
Lambda = 3.0
> Obj. sample 1: {'edge_cost': 25.06505287957105, 'prize_collected': 226.7623826795313, 'objective': -201.69732979996024}
> Obj. sample 2: {'edge_cost': 37.23012326447901, 'prize_collected': 306.3730702170471, 'objective': -269.1429469525681}
> Jaccard dist.: 0.11428571428571428
> Hamming dist.: 0.9559659090909091
Lambda = 3.5
> Obj. sample 1: {'edge_cost': 25.06505287957105, 'prize_collected': 226.7623826795313, 'objective': -201.69732979996024}
> Obj. sample 2: {'edge_cost': 37.23012326447901, 'prize_collected': 306.3730702170471, 'objective': -269.1429469525681}
> Jaccard dist.: 0.11428571428571428
> Hamming dist.: 0.9559659090909091
Lambda = 4.0
> Obj. sample 1: {'edge_cost': 25.06505287957105, 'prize_collected': 226.7623826795313, 'objective': -201.69732979996024}
> Obj. sample 2: {'edge_cost': 37.23012326447901, 'prize_collected': 306.3730702170471, 'objective': -269.1429469525681}
> Jaccard dist.: 0.11428571428571428
> Hamming dist.: 0.9559659090909091
Lambda = 5.0
> Obj. sample 1: {'edge_cost': 25.06505287957105, 'prize_collected': 226.7623826795313, 'objective': -201.69732979996024}
> Obj. sample 2: {'edge_cost': 41.6153825760545, 'prize_collected': 306.3730702170471, 'objective': -264.7576876409926}
> Jaccard dist.: 0.11764705882352941
> Hamming dist.: 0.9573863636363636
Lambda = 10.0
> Obj. sample 1: {'edge_cost': 29.873970790503922, 'prize_collected': 209.45160128136507, 'objective': -179.57763049086114}
> Obj. sample 2: {'edge_cost': 11.773022910393657, 'prize_collected': 205.71245009237012, 'objective': -193.93942718197647}
> Jaccard dist.: 0.2
> Hamming dist.: 0.9772727272727273
Lambda = 15.0
> Obj. sample 1: {'edge_cost': 5.876474987352459, 'prize_collected': 123.35287700024452, 'objective': -117.47640201289207}
> Obj. sample 2: {'edge_cost': 12.990982215735158, 'prize_collected': 182.45966655838677, 'objective': -169.4686843426516}
> Jaccard dist.: 0.0
> Hamming dist.: 0.9815340909090909
Lambda = 20.0
> Obj. sample 1: {'edge_cost': 1.5234903706826128, 'prize_collected': 88.96918230765598, 'objective': -87.44569193697338}
> Obj. sample 2: {'edge_cost': 5.680730602469671, 'prize_collected': 141.21595993438885, 'objective': -135.53522933191917}
> Jaccard dist.: 0.0
> Hamming dist.: 0.9886363636363636
Lambda = 25.0
> Obj. sample 1: {'edge_cost': 1.5234903706826128, 'prize_collected': 88.96918230765598, 'objective': -87.44569193697338}
> Obj. sample 2: {'edge_cost': 2.4332735323262424, 'prize_collected': 96.91390245614741, 'objective': -94.48062892382117}
> Jaccard dist.: 0.0
> Hamming dist.: 0.9914772727272727
Lambda = 30.0
> Obj. sample 1: {'edge_cost': 1.5234903706826128, 'prize_collected': 88.96918230765598, 'objective': -87.44569193697338}
> Obj. sample 2: {'edge_cost': 2.4332735323262424, 'prize_collected': 96.91390245614741, 'objective': -94.48062892382117}
> Jaccard dist.: 0.0
> Hamming dist.: 0.9914772727272727
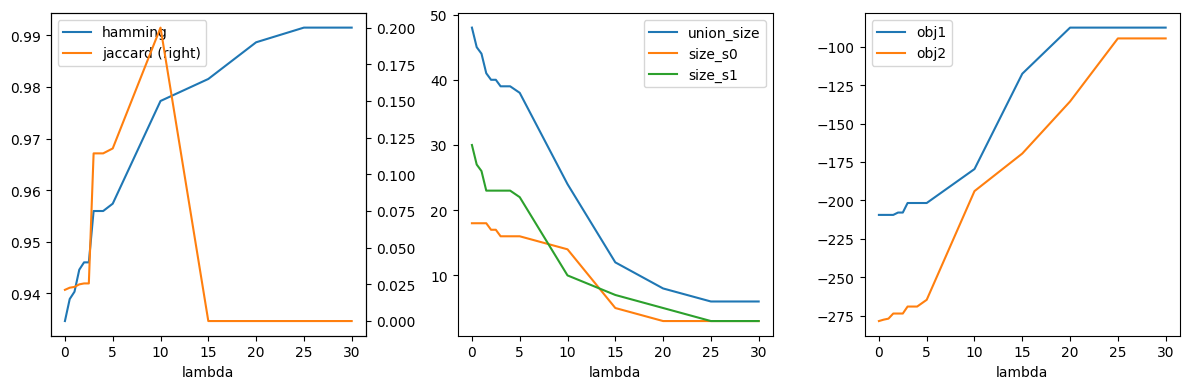
import pandas as pd
fig, axs = plt.subplots(1, 3, figsize=(12, 4))
df_r = pd.DataFrame(
results,
columns=[
"lambda",
"obj1",
"obj2",
"jaccard",
"hamming",
"union_size",
"size_s0",
"size_s1",
],
)
df_r["total_obj"] = df_r["obj1"] + df_r["obj2"]
df_r.plot(x="lambda", y="hamming", ax=axs[0])
df_r.plot(x="lambda", y="jaccard", secondary_y=True, ax=axs[0])
df_r.plot(x="lambda", y=["union_size", "size_s0", "size_s1"], ax=axs[1])
df_r.plot(x="lambda", y=["obj1", "obj2"], ax=axs[2])
plt.tight_layout()
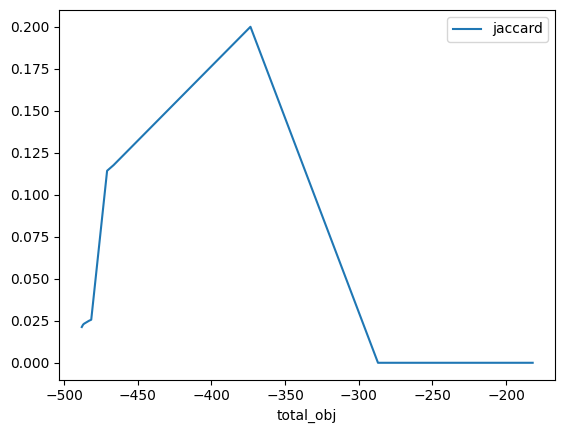
# Compare total objective for the two samples vs jaccard distance
# NOTE: Lower values are better, since the problem is framed as a minimization
df_r.plot(x="total_obj", y="jaccard")
<Axes: xlabel='total_obj'>